pytc¶
Open source python software for extracting thermodynamic information from Isothermal Titration Calorimetry experiments.
- We welcome bug reports, patches, and new thermodynamic models. See the contributing page for how to get involved.
- If you find pytc useful, please star and/or fork the project on github.
- Citation: Duvvuri H, Wheeler LC, Harms MJ. (2018) “pytc: Open-Source Python Software for Global Analyses of Isothermal Titration Calorimetry Data” Biochemistry. doi://10.1021/acs.biochem.7b01264
Download¶
Windows GUI | Mac GUI | Linux GUI | API | Source code
Philosophy¶
- Open source and cross platform. The full source code should be available. The program should not require proprietary software to run.
- Rigorous. Program should use best practices and current algorithms for performing fits and assessing fit quality.
- Ease of use. Fitting basic models should be easy. Implementing completely new thermodynamic models should be straightforward.
- Accessible for users and programmers. It should have both a graphical user interface and a well-designed API.
Features¶
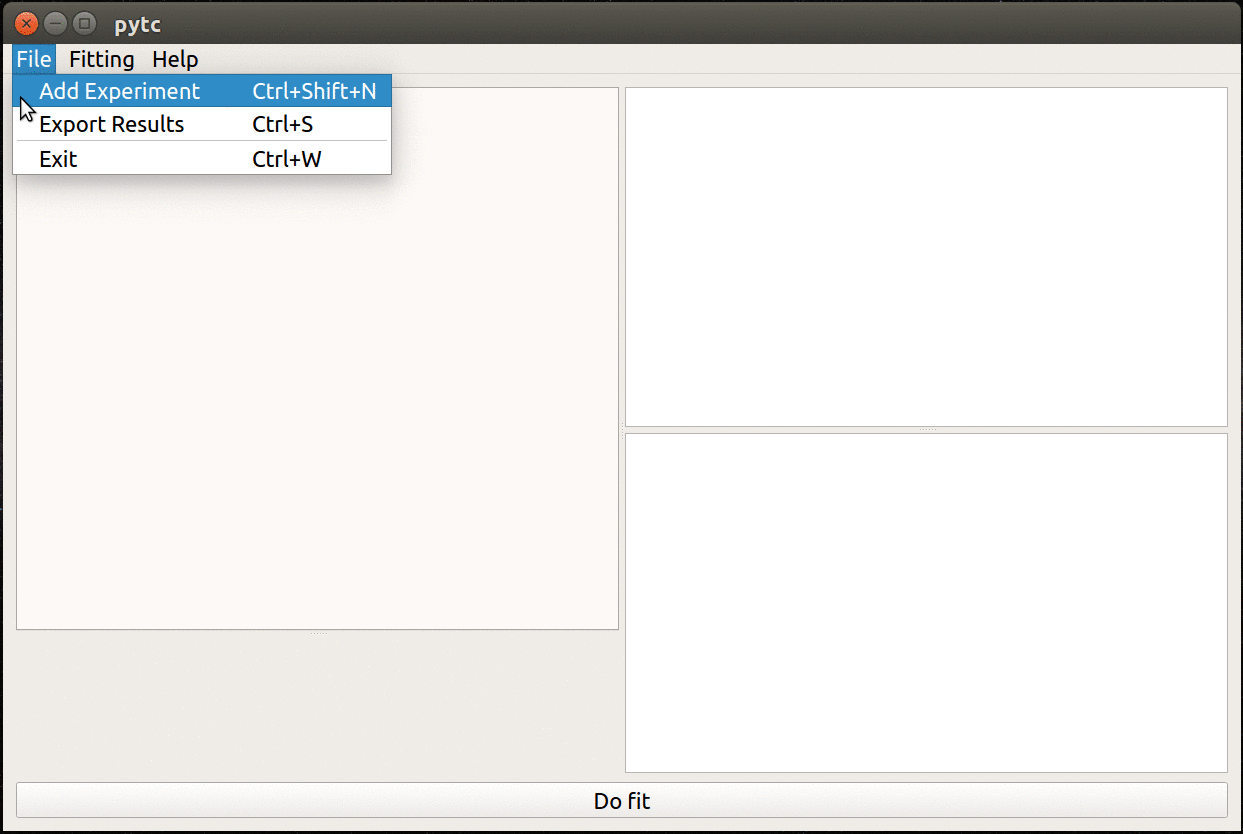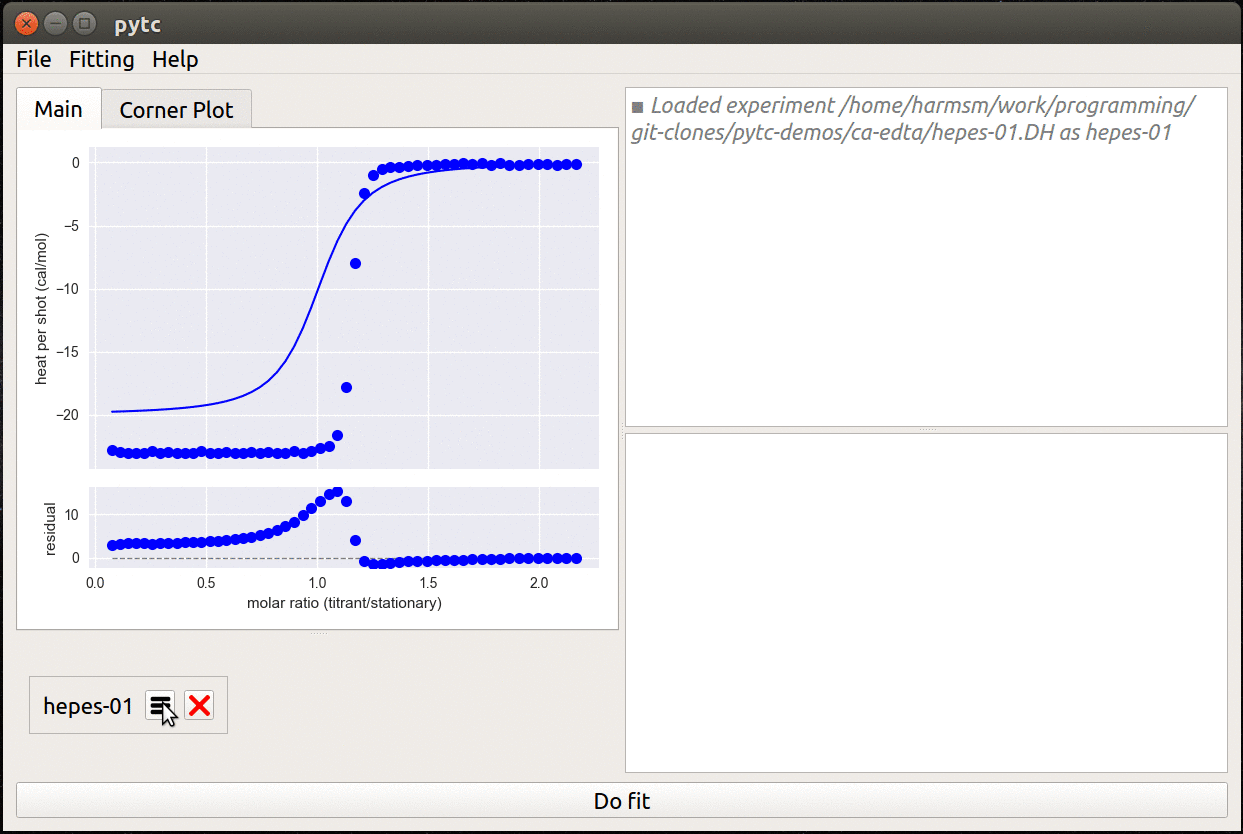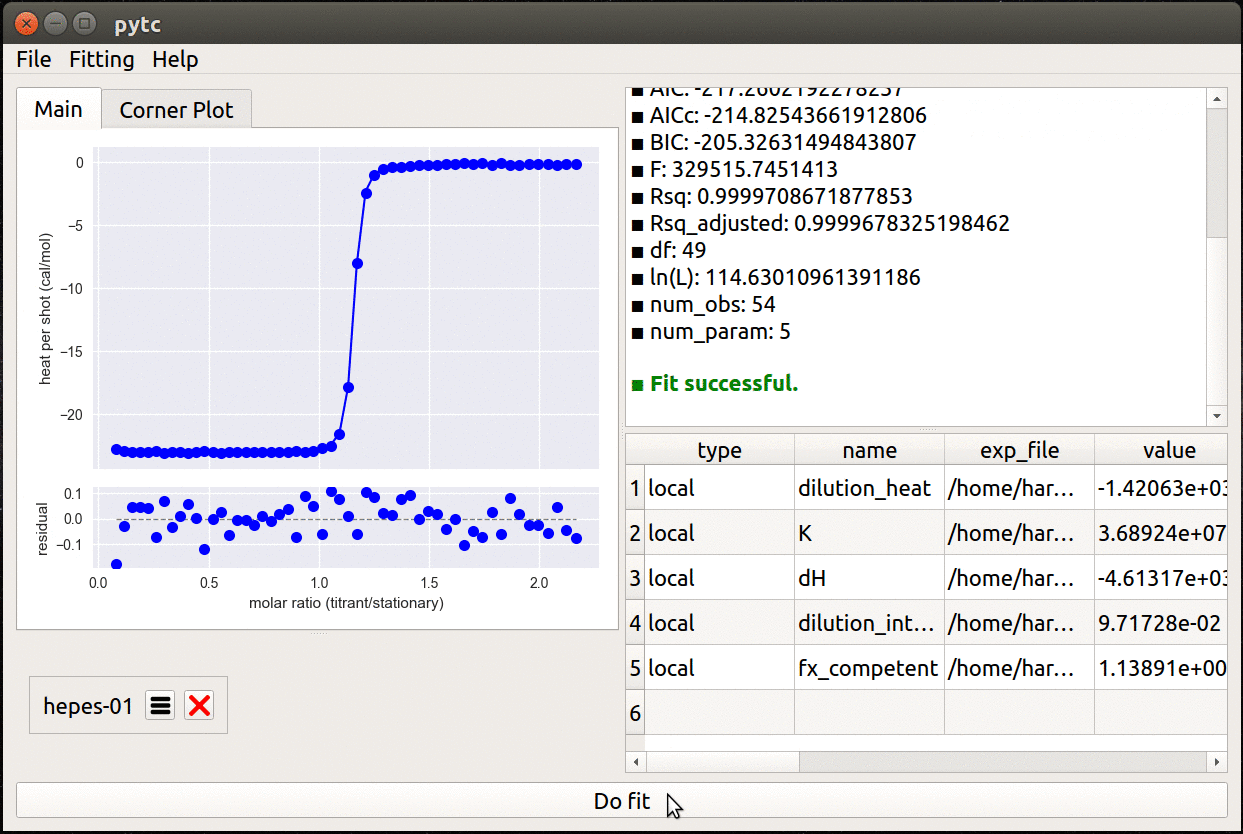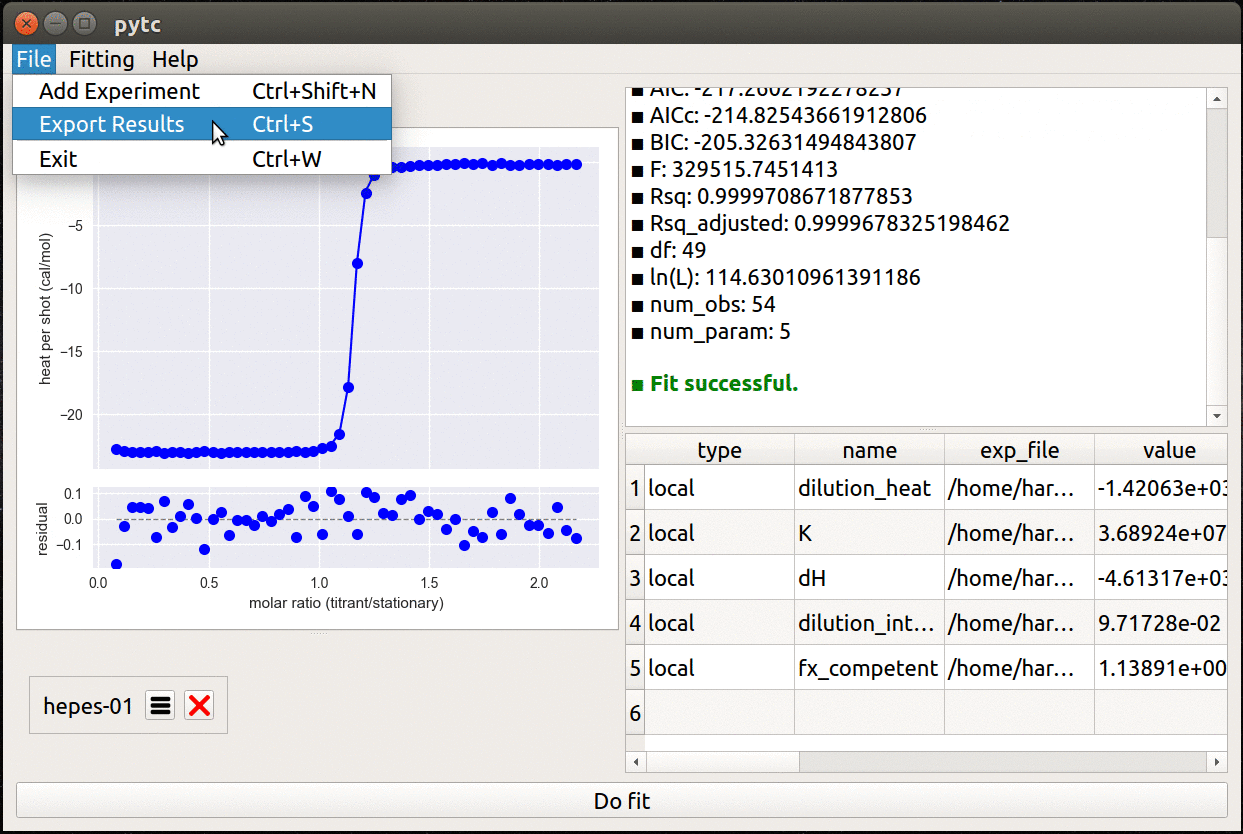
- Simple, cross-platform graphical user interface.
- Clean, pythonic API
- New models can be defined using a few lines of python code
- Easy integration with jupyter notebooks for writing custom fitting scripts
Documentation¶
Warning
pytc will fit all sorts of complicated models to your data. It is up to you to make sure the fit is justified by the data. See the Fitting and statistics section to see what pytc reports to help in this decision making.
API Demo¶
Fit a single-site binding model to a \(Ca^{2+}/EDTA\) binding experiment.
import pytc
# Load in integrated heats from an ITC experiment
e = pytc.ITCExperiment("demos/ca-edta/tris-01.DH",
pytc.indiv_models.SingleSite)
# Create the global fitter, add the experiment
g = pytc.GlobalFit()
g.add_experiment(e)
# Do the fit
g.fit()
# Print the results out
g.plot()
print(g.fit_as_csv)